Öffnet in neuem Fenster
Opens in a new window
Öffnet externe Seite
Opens an external site
Öffnet externe Seite in neuem Fenster
Opens an external site in a new window
How does BESSY II contribute to the research on SARS-CoV-2?
Research on SARS-CoV-2 at BESSY II
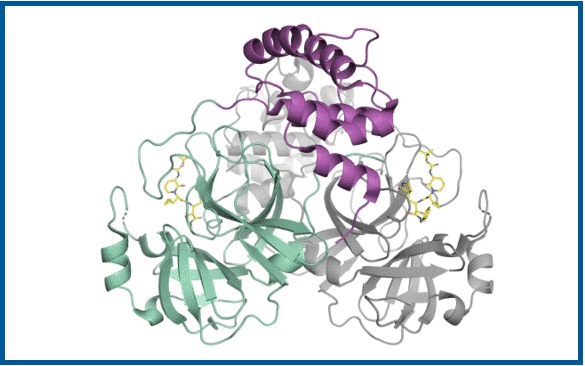
Schematic picture of the coronavirus protease.
(© H. Tabermann / HZB)
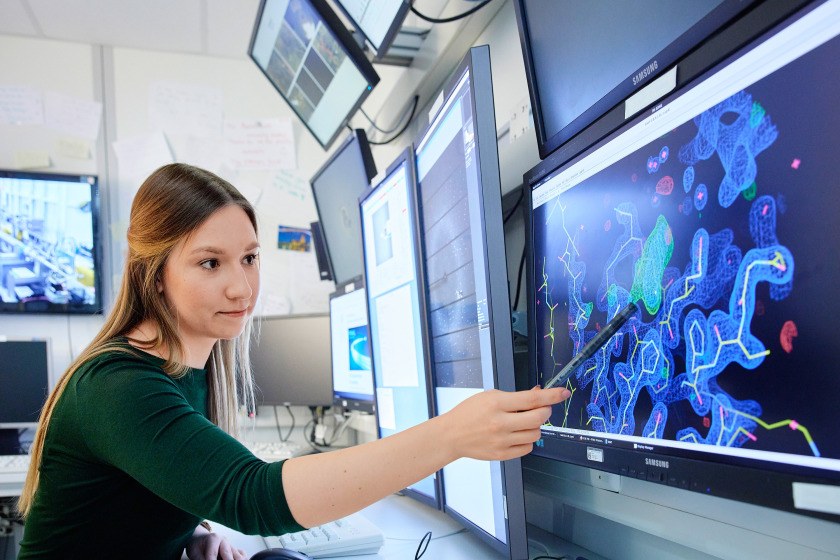
At synchrotron light sources like BESSY II, research is currently gaining crucial insights into combating the SARS-CoV2 virus. The results are helping to contain the spread and fight the disease more effectively.
For corona research, BESSY II has provided access via a fast-track method even during the strictest lockdown phases. Immediately after the genome of the novel coronavirus SARS-CoV2 was sequenced in early 2020, the first measurements of viral proteins started at BESSY II.
- A first major success at the beginning of 2020 was the decoding of the three-dimensional structure of the main protease of the SARS-CoV2 virus, which was already achieved at BESSY II in February 2020. This protein is elementary in the life cycle of the coronavirus because it is involved in the reproduction of the viruses. Knowledge of its 3D structure helps in the search for suitable active substances that dock onto the protein and hinder its function. Because without information about the target protein, the search for an active agent is like looking for a needle in a haystack. Structure-based drug discovery" helps to identify the best candidates for active substances from the multitude of possible substances. > Read more here (news piece)
- The BMBF is currently funding the two projects "CTS-COV-2" and "STOP CORONA" at the two light sources PETRA III and BESSY II. In both projects, the main protease of the virus, which was decoded at BESSY II, was selected as the target for a drug.
- In the STOP-CORONA project, which began as a collaboration between the Helmholtz-Zentrum Berlin (HZB), the University of Lübeck and the University of Würzburg, the aim is to use small organic substances, so-called fragments, to identify active surfaces of the main viral protease. For this fragment screening, the HZB has two libraries available F2X-Entry with 96 substances and F2X-Universal with 1103 substances.
In a first step, crystals of the main protease were tested against the F2X-Entry library. From the binders obtained, a more strongly binding subsequent substance could be identified by optimisation. This substance is currently in binding studies and will be further optimised.
These results provide important insights for drug discovery against SARS-CoV-2, as drugs are still urgently needed to get COVID19 under control. However, Corona research at synchrotrons is not limited to X-ray structure analysis.
Other experiments are currently in preparation or being evaluated:
- A team from the Max-Planck Institute for Chemistry is using microspectroscopy at BESSY II to investigate the emission of aerosol particles during various activities such as speaking, singing or coughing.
- Using the X-ray microscope at BESSY II, a team from Freie Universität Berlin (FU) and HZB led by Prof. Dr. Gerd Schneider has succeeded in spatially imaging coronaviruses in cells. They are currently investigating how virus uptake is blocked by clinically approved drugs from the antidepressant group.
BESSY II research (media and events)
30.06.2021 Synchrotron Radiation News (vol 34)
Keeping science going at BESSY II
This article describes how the teams of BESSY II synchrotron have been working during the pandemic and how they have committed to keep the research going.
Synchrotron Radiation News has created a series in which light sources around the world share their experiences of the pandemic: "How Has COVID-19 Affected Operations and Research? More >
29.06.2021 German Committee Research with Synchrotron Radiation (KFS)
Synchrotrons accelerate corona research
The German Committee Research with Synchrotron Radiation (KFS) has recently published scientific results about corona, BESSY II is mentioned.
10.11.2020 Berlin Science Week From Protein to medecine (hzb youtube channel)
Proteins are the building blocks of life. In his talk Manfred Weiss (group leader of the joint research group macromolecular crystallography) explains how protein crystallography can help in the development of active substances against diseases.
26.6.2020 Helmholtz Association:
Using X-ray imagines to study the virus
Medical and biochemical laboratories are not the only places where research on the new coronavirus is taking place. Particle accelerators are also contributing to decoding SARS-CoV-2 and identifying promising substances for effective medications.
20.05.2020 Mittagsmagazin Das Erste (national TV broadcast, in german): Die Suche nach dem Corona-Medikament
The search for a drug against COVID-19
15.05.2020 Article in nature: The sprint to solve coronavirus protein structures - and disarm them with drugs
Stopping the pandemic could rely on breakneck efforts to visualize SARS-CoV-2 proteins and use them to design drugs and vaccines.
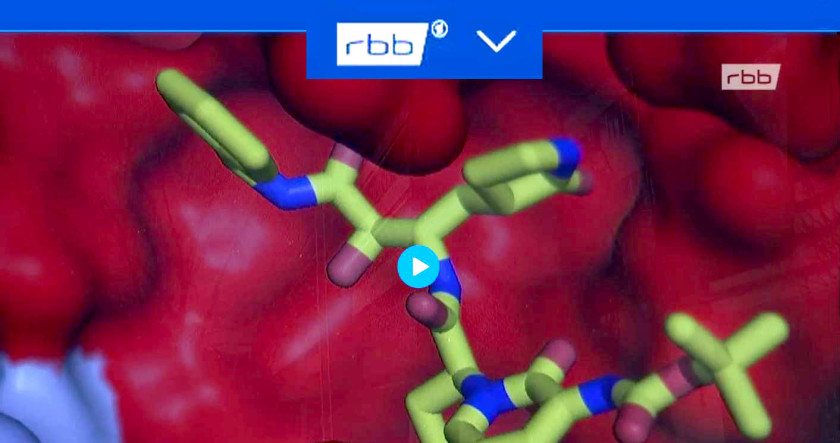
4.05.2020 - Regional broadcast (rbb)
Researchers use the particle accelerator BESSY II to search for an active substance.
Video available online until 11.05.2020
14.04.2020 - Radio broadcast (Deutschlandfunk, in german): Shutdown für Teilchenbeschleuniger
Shutdown for particle accelerators
1.04.2020 - Regional broadcast (rbb, in german): BESSY II research related on SARS-CoV-2
Online until 8.04.2020
23.03.2020 Helmholtz Association: Blueprint for an active substance against the novel coronavirus
Researchers working at the BESSY II X-ray source have decoded the three-dimensional structure of one of the major proteins of the novel coronavirus. Manfred Weiss from the Helmholtz-Zentrum Berlin explains how this could speed up the search for a drug to treat the disease caused by the virus.
20.03.2020 Welt der Physik (article in german): Das erleichtert die Suche nach passenden Wirkstoffen
"This facilitates the search for suitable active substance" – interview with Manfred Weiss (group leader at the Joint Research Group Macromolecular Crystallography at BESSY II)
13.03.2020 - TV broadcast NDR (in german) : Neuer Wirkstoff aus Lübeck soll gegen Coronavirus helfen
New substance from Lübeck to help against coronavirus
Networks on SARS-CoV-2 research
Researchers all over the world are working on active substances against the corona virus. The HZB is part of the following research initiatives:
As Germany's largest research organization, the Helmholtz Association makes important contributions to overcoming the corona crisis through top-level research, please find here their SARS-CoV-2 coronavirus topic page.
The European operators of synchrotron sources and free electron lasers (FEL) have joined forces in the strategic initiative called LEAPS (League of European Accelerator-based Photon Sources). Mid-May LEAPS has published the brochure: Research at LEAPS facilities fighting COVID-19
BESSY II is also member of the international collaboration Ligthsources.org, where communicators from synchrontons and FELs put all their effort together creating “one voice for brightest science”. They collate all the results on COVID-19 related research here: Lightsource research on SARS-CoV-2.
Latest news about research at BESSY II
Please find here the latest news on research done at BESSY II related to SARS-CoV-2.
Scientific publications (BESSY II research)
Linlin Zhang; Daizong Lin; Xinyuanyuan Sun; Ute Curth; Christian Drosten; Lucie Sauerhering; Stephan Becker; Katharina Rox; Rolf Hilgenfeld:
Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors,
Science 20 Mar 2020: eabb3405
doi:
10.1126/science.abb3405
Lei, J.; Hansen, G.; Nitsche, C.; Klein, C.D.; Zhang, L.; Hilgenfeld, R.:
Crystal structure of Zika virus NS2B-NS3 protease in complex with a boronate inhibitor , Science 353 (2016), p. 503-505
doi:
10.1126/science.aag2419
Lei, J.; Hilgenfeld, R.:
Structural and mutational analysis of the interaction between the Middle-East respiratory syndrome coronavirus (MERS-CoV) papain-like protease and human ubiquitin. , Virologica Sinica 31 (2016), p. 288-299
doi:
10.1007/s12250-016-3742-4